ORIGINAL ARTICLE
SILVA, Maria Goretti Varejão da [1], ANDRADE, Jéssica Martins de [2], MOURA, Fernanda Maria de Lino [3], MEDEIROS, Anna Karolyne de Araujo [4], CORDEIRO, Geovania de Souza [5], MELO, Nataly Sayonara da Silva [6], ROLIM, Maria Betânia Queiroz [7], MOURA, Vilton Edson Figueirôa de [8], SILVA, Daniel Dias da [9], SOARES, Anísio Francisco [10], MEDEIROS, Elizabeth Sampaio de [11]
SILVA, Maria Goretti Varejão da. et al. Enterococcus spp. resistant to antimicrobials and biofilm formers in coalho cheese. Revista Científica Multidisciplinar Núcleo do Conhecimento. Year. 08, Ed. 06, Vol. 01, pp. 05-31. June 2023. ISSN: 2448-0959, Access link: https://www.nucleodoconhecimento.com.br/biology/resistant-to-antimicrobials, DOI: 10.32749/nucleodoconhecimento.com.br/biology/resistant-to-antimicrobials
ABSTRACT
The objective of this study was to accomplish a literature review on antimicrobial-resistant Enterococcus spp. and biofilm formers strains in coalho cheese originating from the Brazilian Northeast. The impact of the presence of enterococcal bacteria forming biofilm on coalho cheese and its impacts on public health for consumers of this type of food were observed. However, more microbiological studies on coalho cheese should be conducted due to its importance for the economy and public health.
Keywords: Enterococcus spp, Microorganisms, Cheeses, Drug Resistance, Public Health.
INTRODUCTION
The coalho cheese is a typical Brazilian food, present in the Northeast region for over 150 years and widely consumed in this region, it is made from raw or pasteurized milk. States like Pernambuco, Ceará, Rio Grande do Norte and Paraíba are the main producers of this type of cheese. Its importance in the economy is considerable and significant for milk suppliers, especially those who don’t have access to milk processing units (SILVA et al., 2012).
Enterococci, Gram-positive bacteria isolated from soil, surface water and seawater, also in association with plants, are present in fermented food products. They are Lactic Acid Bacteria (LAB) widely used as starter cultures in the dairy industry for the production of cheese, yogurt and cultured milk (GIRAFFA, 2003; GIRAFFA e ROSSETTI, 2004). Generally, they don’t cause adverse effects in healthy individuals, however, they can also present themselves as opportunistic pathogens related to nosocomial infections, particularly in immunocompromised patients (LOSSOUARN et al., 2019).
Nosocomial infections, caused by intra-hospital microbiota and sometimes conditioned by the healthcare personnel’s microbiota and the patient’s own, are a problem of great clinical and epidemiological importance. Due to causing high rates of morbidity and mortality, as well as extended hospitalization days and economic resource waste, they are considered an emerging situation worldwide (MONTOYA et al., 2015).
In addition, the intrinsic resistance of Enterococcus spp. to various antimicrobials and the acquisition of resistance to others, such as vancomycin, often used in the treatment of patients with severe infections caused by Gram-positive bacteria, has led to the emergence of E. faecalis as a relevant nosocomial pathogen (ANDRADE, 2018).
Enterococcus spp. genus is described as the cause of at least 10% of hospital infections, according to Hollenbeck and Rice (2012), and the main species associated with these infections, E. faecalis (85-90%) and E. faecium, have intrinsic resistance to various antimicrobials such as β-lactams (penicillin and cephalosporins), aminoglycosides (streptomycin), lincosamides (clindamycin in E. faecalis), streptogramins (the quinupristin- dalfopristin combination in E. faecalis) and sulfamethoxazole-trimethoprim. The uncontrolled use of antimicrobial drugs has triggered an increase in strains resistant to treatments, and this scenario becomes more concerning when microorganisms present themselves in the form of biofilm. Complex structures of microbial clusters protected by a layer of exopolysaccharides (EPS), biofilms prevent drug penetration and hinder the treatment of infections (COSTERTON et al. 1999; URQUHART et al. 2019).
Despite the aforementioned, there is still a lack of records in literature regarding antimicrobial-resistant Enterococcus spp. and biofilm-forming strains isolated from animal- derived foods. Therefore, researching these microorganisms with such characteristics in animal-derived foods becomes relevant.
METHODOLOGY
The present study describes a qualitative, narrative literature review in which 90 articles were finally selected for inclusion and full reading. The database was the selection of articles available on Google Scholar, Scielo, and PubMed, where 34, 14 and 42 articles respectively were selected for possible final insertion in the review. The selection of articles occurred during 2020 and took into consideration papers published in Spanish, English, and Portuguese. Papers that presented the theme proposed by the present research were selected. During the search of the bibliographic material the keywords were used: enterococcus, enterococcus durans, enterococcus faecalis, enterococcus faecium, queijo, queijo de coalho, queijo tipo coalho, resistência, resistência antimicrobiana, infecção hospitalar, infecção nosocomial, biofilme, formação de biofilme.
The revision was produced based on the reading, analysis, interpretation, and synthesis of information acquired from selected works that were considered to contain relevant information for the topic addressed in the project. Exclusion criteria for the bibliographic material were: published works in which the results did not apply to the objectives of this review and articles whose study topic was unrelated to the topic. The inclusion criteria took into account papers that covered the specificity of the topic in question and articles and dissertations published in Spanish, English and Portuguese. There were no restrictions regarding the year of publication of the studies.
LITERATURE REVIEW
THE IMPORTANCE OF MILK AND CHEESE IN BRAZIL
According to data from the Brazilian Institute of Geography and Statistics (IBGE,2018), milk production in Brazil has a great impact on the global scenario, being the fifth country with the highest production in the world, losing position only to the European Union, United State, India, and China.
Dairy products under sanitary inspection in Brazil captured 24.45 billion liters of milk in 2018, an increase of 0.5% compared to 2017 (IBGE, 2019), being a very important factor, as milk is a product rich in nutritional and energetic constituents, it is considered a complete food and its consumption is recommended for people of all ages. Produced worldwide, milk is widely used in infant feeding and is a first-line food in combating infant mortality in developing countries (LUNA, 2012).
Cheese is a product that can be matured or fresh, obtained by partially or totally isolating the whey from milk, reconstituted milk, or dairy serum, which are coagulated with the help of enzymes, rennet, bacteria, isolated or combined organic acids (BRASIL, 1996).
Due to its sensory and nutritional properties, cheese is a traditional dairy product with great acceptance in the market, presented for consumption in various varieties that differ in type, flavor, color, shape and aroma, seeking to please the multiple palates of its consumers (NOGUEIRA, 2006).
Cheeses are high biological value foods, rich in proteins, calcium, phosphorus, zinc, iodine, selenium, vitamins and trace elements. There are over 1,000 types of cheese worldwide made from different types of milk and differentiated production processes (LÁCTEA BRASIL, 2006).
With technological development, many varieties have emerged at the national level, some of which are regionally expressed such as coalho cheese (BORGES et al., 2003). According to Andrade (2006), the name derives from the coagulation process, first observed due to the action of coagulating enzymes found in the stomachs of herbivorous animals. One of the typical products from the Northeast region, coalho cheese, has significant cultural and socioeconomic value, it is part of daily food consumption and commonly consumed as an appetizer or as a side dish with other foods. Its production is passed down from generation to generation, mostly through artisanal methods, using traditional practical knowledge transmitted within the family environment (SOUZA et al., 2014). When made by hand, its production has some type of adaptation according to the manufacturer’s skills, physical and economic structure. As a result, the cheeses produced have organoleptic characteristics that give them a regional and cultural identity specific to each region (MENEZES, 2013).
The coalho cheese, both artisanal and industrialized, may not have microbiological safety and standardization due to being a product that is often manipulated and elaborated under unsatisfactory hygienic-sanitary conditions (ANDRADE, 2006), being a considered as a food highly predisposed to contamination (SANTANA et al., 2008).
In order to regulate its production, some regulations are available, such as: the Technical Regulation of Identity and Quality of Dairy Products, which establishes the identity standards and minimum quality requirements that coalho cheese must meet in order to be intended for human consumption (FREITAS FILHO et al., 2009). Federal Law nº 13.860 of 07/18/2019 sanctioned by the President of the Republic of Brazil deals with the production and commercialization of handmade cheese, and related dairy production, while ADAGRO Ordinance nº 007 of 01/04/2018 approves the Technical Regulation for Identity and Quality of handmade coalho cheese in Pernambuco (Imprensa Nacional, 2019; DOE Pernambuco, 2018).
The industrialized cheese (type A) follows the MAPA regulations such as Normative Instruction nº 5 of 02/14/2017, which establishes requirements for evaluating equivalence to the Unified System of Agricultural Health Attention related to physical structure, dependence and equipment of small-scale agro-industrial establishments producing animal products. Finally, MAPA Ordinance 368/97 deals with the technical regulation on hygienic- sanitary conditions and Good Manufacturing Practices for food processing/industrializing establishments.
ENTEROCOCCUS SPP.
Enterococcus spp. bacteria are Gram-positive, oval-shaped cocci that form short chains, belonging to the Enterococcaceae family and have 58 described species (PARTE, 2014). They grow facultatively in both oxygen-rich and oxygen-reduced environments. They are extremely resilient microorganisms to different environmental conditions, supporting varying temperatures from 10°C to 45°C, in hypotonic environments up to 6.5% sodium chloride, as well as in hypertonic environments with pH ranging from acidic (pH between 4 and 9,6), and they can withstand up to 60°C for up to 30 minutes (KONEMAN et al., 2014).
The term Enterococcus was first reported by Thiercelin in 1899, when he described commensal bacteria with the ability to become pathogenic. Due to their morphology and some biochemical similarities, enterococci were initially considered to belong to the genus Streptococcus (SOLACHE, 2019). They are facultative anaerobic cocci, forming chains of various lengths. They are commensal inhabitants of the gastrointestinal tract of humans and other mammals with the ability to survive in environments such as hospitals (ARIAS e MURRAY, 2012).
The Enterococcus spp. and its species come from various origins, such as the environment, animal and human sources. These microorganisms are of great importance as they are part of the normal microbiota of animals and humans, having a similar distribution in both (FISCHER e PHILLIPS, 2009), they constitute a complex group of bacteria that play a dual role in food, as while some strains are used as starter cultures in cheeses and canned foods, giving flavor, odor and texture to these products, other strains are related to food spoilage as potential reservoirs of enterococci that harbor virulence and resistance determinants (TERRA et al., 2018).
From fecal contamination, animal skin, water, milking equipment and expansion tanks, Enterococcus spp. can contaminate milk and its derivatives. Enterococcus bacteria are often found in milk and dairy products, especially in cheeses, due to characteristics such as resistance to pasteurization because they are thermoresistant and refrigeration because they are psychrotrophic (PORTO et al., 2016).
Enterococcus spp. are part of the microbiota involved in various fermentative processes in foods, such as those involving milk, meat and vegetables. They are natural components of food and play an important role in cheese ripening and flavor enhancement. These microorganisms have a wide adaptability to withstand adverse conditions such as temperature, pH, hyperosmolarity and prolonged desiccation (ALI et al., 2014).
Its presence in food has been a concern for public health agencies, mainly due to its ambiguous characteristics (MORAES et al., 2012). On the positive side, there is the development of sensory properties through biochemical reactions during maturation: proteolysis, lipolysis, use of citrate and production of volatile aromatic compounds (HUGAS et al., 2003; JAMET et al., 2012), on the other hand, they have become important human pathogens among hospitalized patients as well as elderly individuals with underlying serious illnesses (LECLERQ, 2009; ROSENTHAL et al., 2015; SIEVERT et al., 2013; ARCHAMBAUD et al., 2019). In recent years, it has been associated with bacteremia, sepsis, and bacterial endocarditis (HEIDARI et al., 2017), these infections can originate from the normal microbiota or can also be transferred from patient to patient or acquired through the consumption of contaminated water or food (MURRAY et al., 2004).
Enterococcus spp. were initially classified as group D Streptococcus due to their species having the cell wall antigen of group D associated with their cytoplasmic membrane (TEIXEIRA e MERQUIOR, 2013). Among the more than 50 species of the genus Enterococcus spp., E. faecalis and faecium are most frequently isolated from humans,
animals, and food samples. From 80 to 90% of Enterococcus spp. isolated from clinical samples in humans are of the species E. faecalis, 5 to 15% are E. faecium, but other species may also occasionally be identified in these isolates such as E. hirae, E. durans, E. gallinarum or E. casseliflavus (SILVA et al., 2012), among these, Enterococcus durans is commonly found in dairy products and the intestinal tract of domestic animals, associated with encephalomalacia and diarrhea in animals. However, unlike E. faecalis or E. faecium, E. durans is not a common pathogen in humans (BYUNG et al., 2019).
ANTIMICROBIAL RESISTANCE AND ENTEROCOCCUS SPP.
The emergence of antimicrobial resistance in enterococci and its spread in food suggest a public health risk, and a possible correlation between strains present in hospitals with those isolated from food should be considered (RIBOLDI et al., 2009). The appearance of bacterial multiresistance in the hospital environment has been increasing in recent decades, increasing morbidity and mortality, length of hospital stay, consequently healthcare costs, challenging public health and increasingly raising antibiotic prescriptions (LÍRIO et al., 2019). Among the main hospital bacterial pathogens that are resistant to available antibiotics is the genus Enterococcus, whose infections represent real therapeutic difficulties (BLANCO et al., 2016).
Antimicrobial Resistance (AMR) is growing worldwide and threatens the prevention and cure of infections. In 2016, antimicrobial resistance was responsible for 70.000 deaths and it is estimated that by 2050 it will cause up to 10 million annual deaths. This estimate is based on the current reality of excessive use of antibiotics, incomplete treatments, neglect in infection control, poor sanitation in developing countries and globalization that allows for easy distribution of microorganisms around the world. These factors contribute to AMR, which combined with the few antibiotics in development, result in infections without treatment (BELLO e DINGLE, 2018).
It is known that Enterococcus spp. tend to undergo natural selection with each application of antimicrobial, leading to the formation of an animal reservoir of more resistant Enterococcus spp., which can infect humans both through direct contact with animals and through the ingestion of animal-derived food (KASZANYITZKY et al., 2007; OLIVEIRA et al., 2010; VIGNAROLI et al., 2011).
Genetic resistance to harmful substances is a natural protective process in some bacterial species that, in some cases, was predetermined millions of years ago (BARLOW e HALL, 2002; LEBRETON et al., 2017). However, the excessive and indiscriminate use of antibiotics in both human infection treatment and livestock farming has resulted in an exponential increase in resistance rates. Currently, bacterial resistance is considered one of the biggest problems related to public health associated with morbidity and mortality (OLIVEIRA, 2019).
It is suggested that the use of antimicrobials in animal feed as growth promoters has created large reservoirs of transferable antibiotic resistance genes in various ecosystems and, consequently, a possible route for transmission of resistant Enterococcus spp. via the food chain (SHEPARD e GILMORE, 2002).
The Enterococcus spp. genus has clinical relevance based not only on its growing prevalence in recent decades, but also on the high rate of antibiotic-resistant strains (MEDEIROS, 2011). Riboldi et al. (2009), when analyzing “natural” foods and dairy products obtained in Porto Alegre/RS, Brazil, observed a high frequency of resistance to antimicrobials used in agriculture, also found an incidence of resistance to nitrofurantoin, an antimicrobial used in the treatment of genitourinary infections, in isolates from cheese and cabbage.
Despite revolutionizing medical practices and being among the greatest advances of the last century, antimicrobial agents have a limited period of clinical usefulness due to the emergence of resistance (LUO et al., 2019).
The increase in antimicrobial resistance is a growing problem, not only in the hospital environment but also for non-hospitalized patients (PÓVOA et al., 2019), as it greatly affects the prevention and satisfactory treatment of a significant number of infections caused by bacteria, parasites, viruses and fungi (CAMACHO et al., 2018).
The improper use of antimicrobials can worsen an infection and lead to the development of bacterial resistance in its treatment (OLIVEIRA e OLIVEIRA, 2014). Having a deeper reflection on the subject, it is known that there is natural resistance, which is important, but the human attitude of using antimicrobials rationally can result in a reversal of this aspect of resistance, gaining mainly public health (CAMACHO et al., 2018).
One of the biggest public health problems, which is antimicrobial resistance, has high clinical relevance because it hinders the control of infectious diseases when therapeutic efficacy decreases, leading to an increase in morbidity and mortality rates. Not to mention that it brings high costs in healthcare and can also promote the transmission of infections to other individuals (MONTEMAYOR et al., 2014). In hospital services, especially in developing countries, there is a serious health problem which is the emergence of bacterial resistance to antibiotics (PEREIRA et al., 2013).
In the health field, since 1944 when the first cases of bacterial resistance were identified, it has been possible to observe the rapid evolution of this episode. After 38 years since the creation of vancomycin for clinical use in Europe in 1988, resistant enterococci to this antimicrobial were detected. In the United States, in 2002, Staphylococcus aureus had the same resistance profile as enterococci, with the aggravating factor being the use of vancomycin as the antibiotic of choice for treating Staphylococcus aureus in most infections caused by this microorganism (DERDE et al., 2012). In Brazil, the panorama of bacterial resistance is concerning, and the increasing emergence of new strains of multidrug-resistant bacteria in hospitals is causing concern among healthcare professionals (COSTA, 2013).
The resistance mechanisms of many microorganisms to antibiotics are variable, with diverse physiopathological aspects such as those of enzymes like β-lactamases that destroy penicillins and some cephalosporins, others modifying enzymes inactivate chloramphenicol and aminoglycosides such as streptomycin and gentamicin. Another mechanism of resistance is directed towards the antibiotic transport form such as resistance to tetracyclines, chloramphenicol, and fluoroquinolones. A third type of mechanism alters the drug intracellularly, for example, the ribosome, metabolic enzymes, or proteins involved in Deoxyribonucleic Acid (DNA) replication or cell wall synthesis, causing the antibiotic to not inhibit a vital function in the microbial cell. More than one type of mechanism can provide resistance to the same antibiotic (NIKAIDO, 1996; LEVY e MARSHALL, 2004).
The growth of Vancomycin-Resistant Enterococcus spp. (VRE) in hospital environments is a major concern, as VRE has the ability to transmit its resistance to other microorganisms, including Methicillin-Resistant Staphylococcus aureus. (MRSA), which is why it is closely monitored and investigated in hospital units (FARON et al., 2016).
According to Terra et al. (2018), antimicrobial resistance in Enterococcus spp. mainly occurs through alterations in the antimicrobial target site and can occur through different mechanisms: alteration of the cell wall, alteration of the cell membrane, alteration of protein synthesis, inhibition of DNA synthesis, competitive inhibition of folic acid synthesis and inhibition of Ribonucleic Acid (RNA) synthesis.
According to the Center for Disease Control and Prevention CDC (2019), for many years, the only therapeutic option to treat infections caused by methicillin-resistant Staphylococcus aureus was vancomycin. The vancomycin resistance gene mediated by plasmid, vanA, is regularly found in vancomycin-resistant isolates associated with medical care for enterococci. Therefore, there is a great opportunity for the transfer of vancomycin resistance genes to S. aureus in patients or locations where the organisms coexist. In the last
15 years, new drugs such as daptomycin, linezolid, and oritavancin have arrived for the treatment of Gram-positive infection resistance. Although resistance to these has been confirmed in some cases, they have considerably high activity against methicillin-resistant Staphylococcus aureus and have the potential to treat vancomycin-resistant S. aureus.
Standardized systems for monitoring the use of antibiotics are an essential requirement as part of an antibiotic resistance control strategy. The use of antibiotics can be quantified in a hospital environment to understand the costs, sensitivity, and routine antibiotic susceptibility in each healthcare service (TERÁN et al., 2018). Three important aspects for success in combating antimicrobial resistance, according to the literature, would be optimizing the administration of existing antimicrobial agents, preventing transmission of drug-resistant organisms through infection control, and reducing environmental contamination (DAVEY et al., 2017).
NOSOCOMIAL INFECTION AND ENTEROCOCCUS SPP.
According to Brazilian Ministry of Health (MS) Ordinance No. 2616/1998 (BRAZIL, 1998), nosocomial infection is one acquired after patient admission and that manifests during hospitalization or after discharge when it can be related to hospitalization or procedures. According to Monteiro (1993), this infection can be defined as any infection related to hospitalization in healthcare facilities, where the patient did not have the pathology during incubation upon admission to the hospital, unless there was a prior hospitalization related to it.
Hospital Infections (HIs) are a public health problem due to the morbidity and mortality they can cause, as well as the social and economic costs associated with them. They can be defined as diseases that affect patients during hospitalization or after discharge. The Center for Disease Control and Prevention (CDC) emphasizes that the infection should not be present during the incubation period while hospitalized or related to previous hospitalizations (COSTA et al., 2019).
The presence of genetic determinants that encode mechanisms of resistance to antimicrobials used in medical clinics is the basis for the ability to colonize and persist in hospital environments (TERRA et al., 2017). Even though they are part of the gastrointestinal tract, Enterococcus spp. are considered opportunistic agents that frequently cause hospital infections.
Among the species that cause the most pathogenesis are Enterococcus faecalis with a rate of 90% and Enterococcus faecium with 10% (KRAHN e FACHINETTO, 2019). Enterococci are considered opportunistic pathogens that often cause infections in patients hospitalized for a long period of time and/or treated with various antibiotics (REIS et al., 2001), and these infections can be in the gastrointestinal tract, urinary tract, skin lesions, heart, and blood (KAO e KLINE, 2019).
Enterococcus spp. species have the ability to adapt widely, facilitating their colonization in hospital environments, which can lead to patient contamination (LEBRETON et al., 2014). As an opportunistic microorganism, it is concerning that in recent years studies have shown a gradual increase in the detection of virulence factors in Enterococcus spp. strains isolated from clinical sources, which has also led to the observation of these virulence genes in strains isolated from food samples (WHO, 2017).
According to Teixeira (2019), the formation of biofilm on hospital devices, as a consequence of microorganisms ability to adhere to surfaces, has led to an increasing development of nosocomial infections that represent a public health problem. Also, according to Fleming et al. (2016) and Omar et al. (2017), antimicrobial resistance may possibly increase in biofilms as a result of the spread of resistance genes among cells through horizontal gene transfer facilitated by the proximity of biofilm cells.
BIOFILM – FORMATION
Biofilm is a structure composed of an aggregate of microorganisms surrounded by an extracellular matrix of polymeric exopolysaccharides that adhere to a solid surface (SHIRTLIFF et al., 2002; COSTERTON et al., 1999). The term biofilm emerged to describe the sessile microbial life form, defined by the adhesion of microorganisms to solid surfaces, with consequent production of extracellular polymeric substances, forming a gelatinous network that immobilizes and protects cells. The formation of biofilms causes phenotypic alterations in planktonic cells, which can be described as survival strategies of microorganisms in environments with adverse conditions (COSTERTON et al. 1999).
Microorganisms in natural environments that exist in the form of biofilms can be detected in the vast majority of substrates that have sufficient moisture to support their growth, with a detection rate between 95% and 99%. Regarding the composition of the biofilm, it is observed that water is the most significant fraction, reaching up to 97% of the biofilm matrix (OLIVEIRA et al., 2010). They are intensely studied not only for their effects on health and the environment but also because they have enormous potential as tools for biotechnological processes (HANSEN et al., 2019).
The ability to form biofilm is an important virulence characteristic of bacteria (DONELLI e GUAGLIANONE, 2004). Thus, the detection of biofilm-producing strains is of great importance for establishing control policies, since failures in the cleaning process allow residues adhered to equipment and surfaces to become a potential source of contamination in the food industry and hospital environments (CASSENEGO et al., 2013).
The formation of bacterial biofilm can be divided into four stages. First, there is the initial attachment of bacterial cells, followed by aggregation and formation of multiple layers of cells, then maturation of the biofilm, and finally detachment of cells from the biofilm to initiate a new cycle of biofilm formation elsewhere (Figure 01). The initial interactions that occur between bacteria and the surface are nonspecific and driven by different forces: hydrophobic, electrostatic, and Lifshitz-Van Der Waals forces, among others (GUTIÉRREZ et al., 2012; SÁNCHEZ et al., 2014).
With the onset of cellular maturation, cells begin to divide and form small cell aggregates characterized by the occurrence of Quorum Sensing (QS) (FONSECA, 2011), which is a communication mechanism between bacteria through the production and diffusion of small chemical or signaling molecules. QS allows coordination of bacterial behavior in relation to the environment by regulating expression of specialized genes in response to population density (LAZAR, 2011; PINTO e FONSECA, 2018).
Figure 01. The four stages of bacterial biofilm formation on surfaces
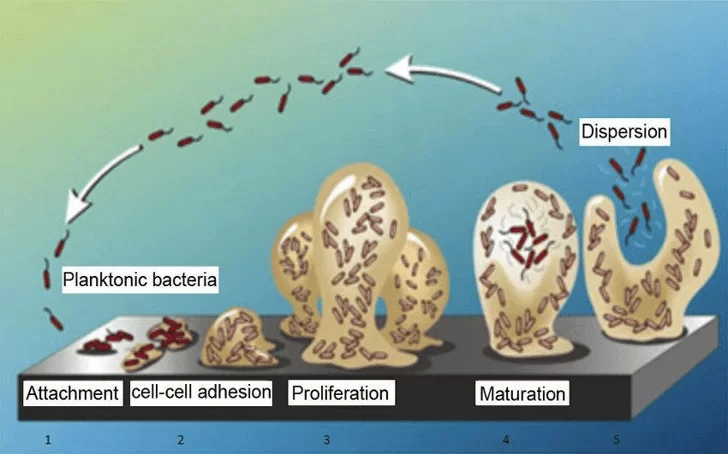
According to Costa et al. (2018), the dairy processing food industry has a major problem with biofilm formation, which is usually associated with serious hygiene problems that can cause product deterioration and recontamination. On the other hand, according to Bosman et al. (2014), one of the main concerns of modern medicine is biofilm-associated infection within the hospital environment, as it has increased its incidence due to the increase in use of implantable medical devices, which have directly contributed to high rates of morbidity and mortality in hospitalized individuals.
FORMATION OF BIOFILM BY ENTEROCOCCUS SPP.
Enterococcus spp. associated with foodborne outbreaks have virulence factors, such as the ability to form biofilms. Microbial aggregation by this bacterial species is apparently influenced by various aspects, such as temperature, exposure to nutrients and sanitizers, and surface characteristics (COSTA et al., 2018).
Enterococci isolated from food that demonstrate the ability to form biofilms are alarming, as this capacity contributes to the survival, persistence, and spread of resistant enterococci and/or resistance genes in various environmental conditions (MEDEIROS et al., 2014).
According to Estrela et al. (2009), the phases of structural organization of biofilm, the composition and activities of colonizing microorganisms in various environments may differ, although the establishment of a micro-community on a surface seems to essentially follow the same series of developmental stages.
According to Medeiros (2011), studies have shown that different factors can influence the formation of biofilm in Enterococcus spp., in addition to conditions that generally affect biofilm formation among a variety of microorganisms, such as pH, CO2 concentration, temperature, among others. There is still no consensus on the influence of glucose concentration on biofilm formation by Enterococcus spp., but some researchers believe that glucose increases biofilm production by Enterococcus spp.
Among the species of this genus, Enterococcus faecalis is an opportunistic pathogen that causes most enterococcal infections with the ability to adhere to biotic and abiotic surfaces, promoting the formation of a biofilm that allows it to grow and survive in aggressive environments (CÂNDIDO et al., 2010).
Among the different factors involved in the pathogenesis of these organisms, biofilm formation and antibiotic resistance are the most important. The ability to form biofilm has been attributed to the presence of some virulence genes (SAFFARI et al., 2017), so other mechanisms for infection control are also observed, such as preventing biofilm formation by Enterococcus spp. or inhibiting the action of other virulence factors, which could provide an alternative method of therapy, especially considering that antimicrobial treatment is a challenge for this genus (COMERLATO et al., 2013). The robust biofilm formation and inherent resistance to multiple drugs in enterococci have made it a challenging nosocomial pathogen (SURYALETHA et al., 2019).
CONCLUSIONS
Enterococcus are versatile microorganisms that can be found as contaminants in coalho cheese and cause food poisoning. It has emerged as one of the most prevalent Gram-positive bacteria in nosocomial infections mainly due to multidrug resistance to antimicrobials. The studies suggest serious public health concerns, as these microorganisms may have the ability to transfer antimicrobial resistance genes to other microorganisms present in the human and animal gut microbiota, making it impossible to use the drugs for the standard clinical treatment mentioned in the article. Further studies and analyses of Enterococcus in food and their ability to resist antimicrobial agents are needed to elucidate the mechanisms of resistance transfer among these microorganisms.
REFERENCES
ALI, S. A. et al. Environmental Enterococci: I. Prevalence of virulence, antibiotic resistance and species distribution in poultry and its related environment in Karachi, Pakistan. Letters in Applied Microbiology. v. 58, n. 5, p. 423-432, 2014. Disponível em: <10.1111/lam.12208>. Acesso em: 2 jun. 2023.
ANDRADE, A. A. Estudo do perfil sensorial, físico-químico e aceitação de queijo de coalho produzido no estado do Ceará. 2006. Dissertação (Mestrado em Tecnologia de Alimentos) – Centro de Ciências Agrárias, Universidade Federal do Ceará, Fortaleza. Disponível em: <http://www.repositorio.ufc.br/handle/riufc/17217>. Acesso em: 2 jun. 2023.
ANDRADE, S. D. E. Caracterização fenotípica e genotípica de amostras clínicas e indígenas de Enterococcus isoladas de seres humanos: diversidade, virulência e resistência a drogas antimicrobiana. 2018. Tese (Doutorado em Ciências Biológicas (Microbiologia)) – Universidade Federal de Minas Gerais, Belo Horizonte – MG. Disponível em: <http://hdl.handle.net/1843/BUOS-B76JMH>. Acesso em: 2 jun. 2023.
ARCHAMBAUD, C. et al. Intestinal translocation of enterococci requires a threshold level of enterococcal overgrowth in the lumen. Scientific Reports. v. 9, n. 8926, pp. 1-12, 2019. Disponível em: <https://doi.org/10.1038/s41598-019-45441-3>. Acesso em: 2 jun. 2023.
ARIAS, C. A.; MURRAY, B. E. The rise of the Enterococcus: Beyond vancomycin resistance. Nature Reviews Microbiology. v. 10, n. 4, pp. 266-278, 2012. Disponível em: <10.1038/nrmicro2761>. Acesso em: 6 jun. 2023.
BARLOW, M.; HALL, B. G. Phylogenetic analysis shows that the OXA beta-lactamase genes have been on plasmids for millions of years. Journal of Molecular Evolution. v. 55, n. 3, pp. 314-321, 2002. Disponível em: <10.1007/s00239-002-2328-y>. Acesso em: 6 jun. 2023.
BELLO, A.; DINGLE, T. C. What’s that resistance mechanism? Understanding genetic determinants of gram-negative bacterial resistance. Clinical Microbiology Newsletter. v. 40, n. 20, pp 165-174, 2018. Disponível em: <10.1016/j.clinmicnews.2018.10.001>. Acesso em: 6 jun. 2023.
BLANCO, A. L.; FREITAS, L. M.; FONTANA, C. R. ACT. Avaliação de resistência à Terapia Fotodinâmica em Enterococcus faecalis. Revista de Ciências Farmacêuticas Básica e Aplicada. v. 37, n. 1, pp. 1-165, 2016. Disponível em: <file:///C:/Users/Workstation%20Video%20B/Downloads/618-Article%20Text-1879-2-10-20200221.pdf>. Acesso em: 6 jun. 2023.
BYUNG-HAN, R. et al. Clinical characteristics and treatment outcomes of Enterococcus durans bacteremia: a 20-year experience in a tertiary care hospital. European Journal of Clinical Microbiology & Infectious Diseases. v. 38, n. 9, pp. 1743-1751, 2019. Disponível em: <https://pubmed.ncbi.nlm.nih.gov/31243595/>. Acesso em: 6 jun. 2023.
BORGES, M. F. et al. Microrganismos patogênicos e indicadores em queijo de coalho produzido no Estado do Ceará, Brasil. Boletim do Centro de Pesquisa de Processamento de Alimentos. v. 21, n. 1, pp. 31- 40, 2003. Disponível em: <10.5380/cep.v21i1.1146>. Acesso em: 6 jun. 2023.
BOSMAN, W. M. P. F. et al. Infections of intravascular bare metal stents: a case report and review of literature. European Journal of Vascular and Endovascular Surgery. v. 47, n. 1, pp. 87-99, 2014. Disponível em: < https://pubmed.ncbi.nlm.nih.gov/24239103/>. Acesso em: 6 jun. 2023.
BRASIL. Ministério da Saúde. Agência Nacional de Vigilância Sanitária – ANVISA. Portaria n. 2616 de 12 de maio de 1998. Brasília. Disponível em <https://bvsms.saude.gov.br/bvs/saudelegis/gm/1998/prt2616_12_05_1998.html>. Acesso em 19/07/2019.
BRASIL. Ministério da Agricultura. Secretaria de Defesa Agropecuária. Departamento de Inspeção de Produtos de Origem Animal DIPOA. Portaria n. 146 de 7 de março de 1996. Aprova os Regulamentos Técnicos de Identidade e Qualidade de Produtos Lácteos. Diário Oficial da União, Brasília, 11 de março de 1996.
CAMACHO, J. O. I. et al. Prescripción racional de antibióticos: una conducta urgente. Medicina Interna de México. v. 34, n. 5, pp. 762-770, 2018. Disponível em: <https://pesquisa.bvsalud.org/portal/resource/pt/biblio-984739>. Acesso em: 7 Jun. 2023.
CÂNDIDO, C. S. et al. Effects of Myrcia ovata Cambess. Essential oil on planktonic growth of gastrointestinal microorganisms and biofilm formation of Enterococcus faecalis. Brazilian Journal of Microbiology. v. 41, n. 3, pp. 621-627, 2010. Disponível em: <https://pesquisa.bvsalud.org/portal/resource/pt/lil-549403>. Acesso em: 7 Jun. 2023.
CASSENEGO, A. P. V.; ELLWANGER, J.; PEDRO, A.; AZEVEDO, A.; RIBEIRO, M.L.; FRAZZON, A. P. G. Virulência e formação de biofilme microbiano por Enterococcus faecalis isolados de swabs cloacais de frangos de corte infectados com Eimeria spp. Pesquisa Veterinária Brasileira. v. 33, n. 12, pp. 1433-1440 2013. Disponível em: <https://www.scielo.br/j/pvb/a/PsDXt5fJyrS6gjZ9xCtFqfw/?lang=pt>. Acesso em: 7 Jun. 2023.
CENTERS FOR DISEASE CONTROL AND PREVENTION, C.D.C. (online). Antibiotic resistance threats in the United States. 2019. Disponível em:
<https://www.cdc.gov/drugresistance/pdf/threats-report/2019-ar-threats-report-508.pdf>. Acesso em: 22 jul. 2019.
COMERLATO, C. B. et al. Presence of virulence factors in Enterococcus faecalis and Enterococcus faecium susceptible and resistant to vancomycin. Memória do Instituto Oswaldo Cruz. v. 108, n. 5, pp. 590-595, 2013. Disponível em: <https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3970601/>. Acesso em: 7 Jun. 2023.
COSTA, F. T. Síntese, caracterização e o estudo do efeito de nano-partículas de CoFe2O4-YFe2O3 em bactérias patogênicas. 2013. Dissertação (Mestrado em Ciências de Materiais) – Universidade de Brasília, Brasília. Disponível em: <https://repositorio.unb.br/handle/10482/14397>. Acesso em: 7 Jun. 2023.
COSTA, G. A. et al. Evaluation antibacterial and antibiofilm activity of the antimicrobial peptide P34 against Staphylococcus aureus and Enterococcus faecalis. Anais Academia Brasileira de Ciências. v. 90, n. 1, pp. 1-12, 2018. Disponível em: <https://doi.org/10.1590/0001-3765201820160131>. Acesso em: 7 Jun. 2023.
COSTA, M. C. P. et al. Micro-Organismos isolados a partir de espécimes clínicos de centro cirúrgico. Revista Saúde e Desenvolvimento. v. 13, n. 14, pp. 1-21, 2019. Disponível em: <https://www.revistasuninter.com/revistasaude/index.php/saudeDesenvolvimento/article/view/1013>. Acesso em: 7 Jun. 2023.
COSTERTON, J. W.; STEWART, P. S.; GREENBERG, E. P. Bacterial biofilms: a common cause of persistent infections. Science. v. 284, n. 5418, p. 1318-1322, 1999. Disponível em: <https://www.science.org/doi/10.1126/science.284.5418.1318>. Acesso em: 7 Jun. 2023.
DAVEY, P. et al. Interventions to improve antibiotic prescribing practices for hospital inpatients. Cochrane Database of Systematic Reviews. v. 2, n. 2, pp. 1-326, 2017. Disponível em: <https://doi.org/10.1002/14651858.CD003543.pub4>. Acesso em: 7 Jun. 2023.
DERDE, L. P.; DAUTZENBERG, M. J.; BONTEN, M. J. Chlorhexidine body wasshing to control antimicrobial-resistant bacteria in intensive care units: a systematic review. Intensive Care Medicine. v. 38, n. 6, pp. 931-939, 2012. Disponível em: <10.1007/s00134-012-2542-z>. Acesso em: 7 Jun. 2023.
DONELLI, G.; GUAGLIANONE, E. Emerging role of Enterococcus spp. In catheter related infections: biofilm formation and novel mechanisms of antibiotic resistance. The Journal of Vascular. v. 5, n. 1, pp. 3-9, 2004. Disponível em: <10.1177/112972980400500101>. Acesso em: 7 Jun. 2023.
ESTRELA, C. R. A. et al. model system to study antimicrobial strategies in endodontic biofilms. Journal of Applied Oral Science. v. 17, n. 2, 2009. Disponível em: <https://www.scielo.br/j/jaos/a/rzGrQDfDvY7RJMJkMwfJPTx/?lang=en>. Acesso em: 7 Jun. 2023.
FARON, M. L.; LEDEBOER, N. A.; BUCHAN, B. W. Resistance mechanisms, epidemiology, and approaches to screening for vancomycin-resistant Enterococcus in the health care setting. Journal of Clinical Microbiology. v. 54, n. 10, pp. 2436- 2447, 2016. Disponível em: <10.1128/JCM.00211-16>. Acesso em: 7 Jun. 2023.
FISHER, K.; PHILLIPS, C. The ecology, epidemiology and virulence of Enterococcus. Microbiology. v. 155, n. 6, pp. 1749-1757, 2009. Disponível em: <10.1099/mic.0.026385-0>. Acesso em: 7 Jun. 2023.
FLEMMING, H. C. et al. Biofilms: an emergent form of bacterial life. Nature Reviews Microbiology. v. 14, n. 9, pp. 563-575, 2016. Disponível em: <https://www.nature.com/articles/nrmicro.2016.94>. Acesso em: 7 Jun. 2023.
FONSECA, A. P. Biofilms in wounds: An unsolved problem? Working together to ensure better patient outcomes. EWMA Journal. V. 11, n. 2, p. 10- 23, 2011.
FREITAS FILHO, J. R. et al. Avaliação da qualidade do queijo “coalho” artesanal fabricado em Jucati-PE. Revista Eletrônica de Extensão – Extensio. v. 6, n. 8, p. 35-49, 2009. Disponível em: <https://periodicos.ufsc.br/index.php/extensio/article/view/1807-0221.2009v6n8p35/11446>. Acesso em: 7 Jun. 2023.
GIRAFFA, G. Functionality of enterococci in dairy products. International Journal of Food Microbiology. v. 88, n. 2-3, p. 215-222, 2003. Disponível em: <10.1016/s0168-1605(03)00183-1>. Acesso em: 7 Jun. 2023.
GIRAFFA, G; ROSSETTI, L. Monitoring of the bacterial composition of dairy starter cultures by RAPD-PCR. FEMS Microbiology Letters. v. 237, n. 1, p. 133-138, 2004. Disponível em: <https://doi.org/10.1111/j.1574-6968.2004.tb09688.x>. Acesso em: 7 Jun. 2023.
GUTIÉRREZ, D. et al. Incidence of Sthapylococcus aureus and analysis of associated bacterial communities on food industry surfaces. Applied and Environmental Microbiology. v. 78, n. 24, p. 8547-8554, 2012. Disponível em: <10.1128/AEM.02045-12>. Acesso em: 7 Jun. 2023.
HANSEN, S. H. et al. Machine-assisted cultivation and analysis of biofilms. Scientific Reports. v. 9, n. 8933, pp. 1-10, 2019. Disponível em: <file:///C:/Users/Workstation%20Video%20B/Downloads/s41598-019-45414-6.pdf>. Acesso em: 7 Jun. 2023.
HEIDARI, H. et al. High Incidence of Virulence Factors among Clinical Enterococcus faecalis Isolates In Southwestern Iran. Infect Chemother. v. 49, n. 1, p. 51-56, 2017. Disponível em: <10.3947/ic.2017.49.1.51>. Acesso em: 7 Jun. 2023.
HOLLENBECK, B. L.; RICE, L. B. Intrinsic and acquired resistance mechanisms in Enterococcus. Virulence. v. 3, n. 5, p. 421-433, 2012. Disponível em: <10.4161/viru.21282>. Acesso em: 7 Jun. 2023.
HUGAS, M.; GARRIGA, M. M.; AYMERICH, M. T.; Functionality of enterococci in meat products. International Journal of Food Microbiology. v. 88, n. 2, p. 223-233, 2003. Disponível em: <10.1016/s0168-1605(03)00184-3>. Acesso em: 9 Jun. 2023.
IBGE – INSTITUTO BRASILEIRO DE GEOGRAFIA E ESTATÍSTICA. Censo Agropecuário 2017. Rio de Janeiro: IBGE, 2017. Disponível em: <https://sidra.ibge.gov.br/pesquisa/censo-agropecuario/censo-agropecuario-2017/resultados-definitivos>. Acesso em: 22 jun. 2019.
IBGE – INSTITUTO BRASILEIRO DE GEOGRAFIA E ESTATÍSTICA. Produção da Pecuária Municipal 2018. Rio de Janeiro: IBGE, v. 46, p. 1-8, 2018. Disponível em: <https://biblioteca.ibge.gov.br/visualizacao/periodicos/84/ppm_2018_v46_br_informativo.pdf>. Acesso em: 9 fev. 2020.
JAMET, E. et al. Prevalence and characterization of antibiotic resistant Enterococcus faecalis in French cheeses. Food Microbiology. v. 31, n. 2, p.191-198, 2012. Disponível em: <10.1016/j.fm.2012.03.009>. Acesso em: 7 Jun. 2023.
KAO, P. H. N.; KLINE, K. A. Dr. Jekyll and Mr. Hide: How Enterococcus faecalis Subverts the Host Immune Response to Cause Infection. Journal of Molecular Biology. v. 431, n. 16, p. 2932- 2945, 2019. Disponível em: <10.1016/j.jmb.2019.05.030>. Acesso em: 9 Jun. 2023.
KARAGULER, T. KAHRAMAN, H. TUTER, M. Analyzing effects of ELF electromagnetic fields on removing bacterial biofilm. Biocybernetics and Biomedical Engineering. v. 37, n. 2, p. 336-340, 2017. Disponível em: <10.1016/j.bbe.2016.11.005>. Acesso em: 01 maio 2023.
KASZANYITZKY, E. J.; TENK, M.; GHIDÁN, A.; FEHÉVÁRI, G. Y.; PAPP, M. Antimicrobial susceptibility of enterococci strains isolated from slaughter animals on the data of Hungarian resistance monitoring system from 2001 to 2004. International Journal of food microbiology. v.115, n. 1, p. 119-123, 2007. Disponível em: <10.1016/j.ijfoodmicro.2006.10.004>. Acesso em: 7 Jun. 2023.
KONEMAN, E. W. et al. Enterococcus. In: KONEMAN, Elmer W. (Ed.). Diagnóstico microbiológico. 7. ed. Rio de Janeiro: Guanabara Koogan. p. 589-659, 2014.
KRAHN, C. O.; FACHINETTO, J. M. Enterococcus spp.: Características gerais e considerações médicas, uma revisão. 6º Congresso Internacional em Saúde. 2019. Disponível em: <file:///C:/Users/Workstation%20Video%20B/Downloads/10765-Texto%20do%20artigo-41799-1-10-20190424.pdf>. Acesso em: 7 Jun. 2023.
LÁCTEA BRASIL. Queijo – Alimento nobre e saudável. [S. l.: s. n.], 2006. Disponível em: <https://www.yumpu.com/pt/document/read/13019755/queijo-alimento-nobre-e-saudavel-nobre-e-saudavel-capril-virtual>. Acesso em 30 jul. 2019.
LAZAR, V. Quorum sensing in biofilms – how to destroy the bacterial citadels or their cohesion/power? Anaerobe. v.17, n. 6, p. 280-285, 2011. Disponível em: <10.1016/j.anaerobe.2011.03.023>. Acesso em: 7 Jun. 2023.
LEBRETON, F. et al. Tracing the Enterococci from Paleozoic Origins to the Hospital. Cell. v. 169, n. 5, p. 849-861, 2017. Disponível em: <10.1016/j.cell.2017.04.027>. Acesso em: 7 Jun. 2023.
LEBRETON, F. et al. Enterococcus diversity, origins in nature, and gut colonization/ Enterococci: from commensals to leading causes of drug resistant infection. Boston: Massachusetts Eye and Ear Infirmary. p.1-59, 2014. Disponível em: <https://www.ncbi.nlm.nih.gov/books/NBK190427/>. Acesso em: 7 Jun. 2023.
LECLERCQ, R. Epidemiological and resistance issues in multidrug-resistant staphylococci and enterococci. Clinical Microbiology and Infection. v. 15, n. 3, p. 224-231, 2009. Disponível em: <10.1111/j.1469-0691.2009.02739.x>. Acesso em: 7 Jun. 2023.
LEVY, S.; MARSHALL, B. Antimicrobial resistance worlwide: causes, challenges and responses. Nature medicine supplement. v. 10, n. 12, p. 122-129, 2004. Disponível em: <https://www.nature.com/articles/nm1145>. Acesso em: 9 Jun. 2023.
LÍRIO, M.; ANDRADE, T.; MENDES, A. V.; BARBERINO, M. G. Avaliação da colonização por bactérias multirresistentes em pacientes admitidos via central de regulação do estado em um hospital filantrópico em Salvador, Bahia. Revista de Epidemiologia e Controle de Infecção. v. 9, n. 1, 2019. Disponível em: <https://online.unisc.br/seer/index.php/epidemiologia/article/view/11595>. Acesso em: 9 Jun. 2023.
LOSSOUARN, J. et al. Enterococcus faecalis Countermeasures Defeat a Virulent Picovirinae Bacteriophage. Journal viruses. v. 11, n. 1, p. 48, 2019. Disponível em: <https://doi.org/10.3390/v11010048>. Acesso em: 9 Jun. 2023.
LUNA, R. O. Identificação, perfil de resistência a antimicrobianos e caracterização molecular de Aeromonas spp. isoladas de queijos coalho tipo A, comercializados na cidade de Recife PE. 2012. Dissertação (Mestrado em Ciência Veterinária). UFRPE, Pernambuco – Recife.
LUO, X. et al. A diversity-oriented rhodamine library for wide-spectrum bactericidal agents with low inducible against resistant pathogens. Nature Communications. v. 10, n. 258, 2019. Disponível em: <https://www.nature.com/articles/s41467-018-08241-3>. Acesso em: 9 Jun. 2023.
MEDEIROS, A. W. Avaliação dos fatores de virulência e a capacidade de formação de biofilme in vitro em isolados alimentares e clínicos de Enterococcus spp. e utilização de PCR-RFLP para a identificação de Enterococcus casseliflavus e Enterococcus gallinarum. 2011 Dissertação (Mestrado). Universidade Feevale, Rio Grande do Sul.
MEDEIROS, A. W.; PEREIRA, R. I.; OLIVEIRA, D. V.; MARTINA, P. D.; AZEVEDO, P. A.; VAN DER SAND, S.; FRAZZON, J.; FRAZZON, A. P. G. Molecular detection of virulence factors among food and clinical Enterococcus faecalis strains in South Brazil; Brazilian Journal of Microbiology. vol. 45, n.1, 2014. Disponível em: <http://dx.doi.org/10.1590/S1517- 83822014005000031>. Acesso em 22 Jul. 2019.
MENEZES, S. S. M. “Queijo de coalho-rei do balcão”: Expansão da produção alicerçada pela demanda dos migrantes sertanejos. Habitus. v. 11, n. 2, p. 143-158, 2013. Disponível em: <https://doi.org/10.18224/hab.v11.2.2013.143-158>. Acesso em: 7 Jun. 2023.
MONTEIRO, J. A. INFECÇÕES NOSOCOMIAIS. Alguns Aspectos. ACTA Médica Portuguesa. v. 6, p. 135-140, 1993. Disponível em: <https://silo.tips/download/infecoes-nosocomiais-alguns-aspectos>. Acesso em: 7 Jun. 2023.
MONTEMAYOR, J. C. G.; BOFARULL, A. M.; MOCHALES, F. B. Impacto de los movimientos migratorios en la resistencia bacteriana a los antibióticos. Revista Española de Salud Pública. v. 88, n. 6, 2014. Disponível em: <https://scielo.isciii.es/scielo.php?script=sci_arttext&pid=S1135-57272014000600014>. Acesso em: 7 Jun. 2023.
MONTOYA, R. R. et al. Variables associadas a costos em cuidados intensivos. Revista de la Asociación Mexicana de Medicina Crítica y Terapia Intensiva. v. 29, n. 3, p. 138-144, 2015. Disponível em: <https://www.scielo.org.mx/scielo.php?script=sci_arttext&pid=S0187-84332015000300003>. Acesso em: 7 Jun. 2023.
MORAES, P. M. et al. Bacteriocinogenic and virulence potential of Enterococcus isolates obtained from raw milk and cheese. Journal of Applied Microbiology. v. 113, p. 318-328, 2012. Disponível em: <https://www.locus.ufv.br/bitstream/123456789/13991/1/Moraes_et_al-2012-Journal_of_Applied_Microbiology.pdf>. Acesso em: 9 Jun. 2023.
MURRAY, P. et al. Microbiologia Médica. 4 ed. Rio de Janeiro: Guanabara Koogan, p. 220-223, 2004. Disponível em: <https://eu-ireland-custom-media-prod.s3-eu-west-1.amazonaws.com/Brasil/Downloads/03-12/ESAMPLE-9788535290363.pdf>. Acesso em: 9 Jun. 2023.
NIKAIDO, H. Multidrug efflux pumps of gram-negative bacteria. Journal of bacteriology. v. 178, n. 20, p. 5853-5859, 1996. Disponível em: <10.1128/jb.178.20.5853-5859.1996>. Acesso em: 7 Jun. 2023.
NOGUEIRA, J. G. A embalagem como fator de agregação de valor ao produto: Um estudo do segmento de queijos em Juiz de Fora. 2006. Dissertação (Mestrado Profissional em Sistemas de Gestão) – Centro Tecnológico, Universidade Federal Fluminense, Niterói.
OLIVEIRA, A. F.; OLIVEIRA FILHO, H. Perfil microbiológico e de resistência antimicrobiana no pé diabético infectado. Jornal Vascular Brasileiro. v. 13, n. 4, p. 289-293, 2014. Disponível em: <https://www.scielo.br/j/rcbc/a/rPHTb6ZvfYkgd9rcczXdYjf/?lang=pt>. Acesso em: 7 Jun. 2023.
OLIVEIRA, E. S. Emergência de Enterococcus spp. resistentes à vancomicina na cidade do Natal – RN. 2019. Dissertação (Mestrado em Ciências Biológicas), Universidade Federal do Rio Grande do Norte. Natal – RN.
OLIVEIRA, M. M. M.; BRUGNETA, D. F.; PICCOLI, R. H. Biofilmes microbianos na indústria de alimentos: uma revisão. Revista do Instituto Adolfo Lutz. v. 69, n. 3, p. 277-284, 2010. Disponível em: <http://www.ial.sp.gov.br/resources/insituto-adolfo-lutz/publicacoes/rial/10/rial69_3_completa/1289.pdf>. Acesso em: 7 Jun. 2023.
OLIVEIRA, M. et al. Antimicrobial resistance and in vitro biofilm-forming ability of enterococci from intensive and extensive farming broilers. Poultry Science. v. 89, n. 5, p. 1065- 1069, 2010. Disponível em: <10.3382/ps.2008-00436>. Acesso em: 7 Jun. 2023.
OMAR, A. et al. Microbial Biofilms and Chronic Wounds. Journal Microorganisms. v. 5, n. 1, pp. 1-15, 2017. Disponível em: <10.3390/microorganisms5010009>. Acesso em: 7 Jun. 2023.
PARTE, A. C. LPSN-list of prokaryotic names with standing in nomenclature. Nucleic Acids Research. v. 42, n. 1, p. 613-616, 2014. Disponível em: <10.1093/nar/gkt1111>. Acesso em: 7 Jun. 2023.
PEREIRA, F. G. F. et al. Characterization of infections related to health care in a neonatal intensive care unit. Enfermagem UERJ. v. 21, n. 1, 2013. Disponível em: <https://www.e-publicacoes.uerj.br/index.php/enfermagemuerj/article/view/6370/5899>. Acesso em: 7 Jun. 2023.
PINTO, G.; FONSECA, A. F. Biofilmes e Feridas Crônicas: Um potencial algoritmo. Journal of tissue regeneration & healing. 2018. Disponível em: <https://bdigital.ufp.pt/bitstream/10284/5816/1/PPG_25983.pdf>. Acesso em: 9 Jun. 2023.
PORTO, B. C et al. Determinantes de virulência em Enterococcus endógenos de queijo artesanal. Revista Ciência Agronômica. v. 47, n. 1, p. 69-76, 2016. Disponível em: <http://ccarevista.ufc.br/seer/index.php/ccarevista/article/view/4297>. Acesso em: 9 Jun. 2023.
PÓVOA, A. C. et al. J. R. Evolução da resistência bacteriana em infecção comunitária do trato urinário em idosos. Revista de Epidemiologia e Controle de Infecção. v. 9, n. 1, p. 08-14, 2019. Disponível em: <https://online.unisc.br/seer/index.php/epidemiologia/article/view/10468>. Acesso em: 9 Jun. 2023.
REIS, A. O.; CORDEIRO, J. C. R.; MACHADO, A. M. O.; SADER, H. S. In vitro antimicrobial activity of linezolid tested against vancomycin-resistant enterococci isolated in Brazilian hospitals. Brazilian Journal of Infectious Diseases. v. 5, n. 5, 2001. Disponível em: <https://www.scielo.br/j/bjid/a/rNpQ6tBdzyFHjVLktGDt6SK/?lang=en>. Acesso em: 9 Jun. 2023.
RIBOLDI, G. P.; FRAZZON, J.; AZEVEDO, P. A.; Antimicrobial resistance profile of Enterococcus spp. isolated from food in Southern Brazil. Brazilian Journal of Microbiology. v. 40, p. 125-128, 2009. Disponível em: <https://www.scielo.br/j/bjm/a/fzR4WH7LGmKh3QRwk8fW3HH/?lang=en>. Acesso em: 9 Jun. 2023.
ROSENTHAL, V. D. et al. International nosocomial infection control consortium (INICC) report, data summary of 43 countries for 2007-2012. Device-associated module. American Journal of Infection Control. v. 43, n. 7, pp. 779-781, 2015. Disponível em: <https://pubmed.ncbi.nlm.nih.gov/25179325/>. Acesso em: 9 Jun. 2023.
SAFFARI, F.; DALFARDI, M. F.; MANSOURI, S.; AHMADRAJABI, R. Survey for Correlation between Biofilm Formation and Virulence Determinants in a Collection of Pathogenic and Fecal Enterococcus faecalis Isolates. Infection & chemotherapy. v. 49, n. 3, p. 176-183, 2017. Disponível em: <10.3947/ic.2017.49.3.176>. Acesso em: 9 Jun. 2023.
SÁNCHEZ, D. V.; CABO, M. L.; IBUSQUIZA, P. S.; HERRERA, J. J. R. Biofilm forming abilityand resistance to industrial disinfectants of Staphylococcus aureus isolated from fishery products. Food Control. v. 39, n. 1, p. 8-16, 2014. Disponível em: <https://doi.org/10.1016/j.foodcont.2013.09.029>. Acesso em: 9 Jun. 2023.
SANTANA, R. F. et al. Qualidade microbiológica de queijo coalho comercializado na cidade de Aracaju, SE. Arquivo Brasileiro de Medicina Veterinária e Zootecnia. v. 60, n. 6, p. 1517-1522, 2008. Disponível em: <https://doi.org/10.1590/S0102-09352008000600031>. Acesso em: 9 Jun. 2023.
SHEPARD, B. D.; GILMORE, M. S. Antibiotic-resistant enterococci: the mechanisms and dynamics of drug introduction and resistance. Microbes and Infection. v. 4, n. 2, p. 215-224, 2002. Disponível em: <10.1016/s1286-4579(01)01530-1>. Acesso em: 9 Jun. 2023.
SHIRTLIFF, M. E.; MADER, J. T.; CAMPER, A. K. Molecular interactions in Biofilms. Chemistry & biology. v. 9, n. 8, p. 859-871, 2002. Disponível em: <https://doi.org/10.1016/S1074-5521(02)00198-9>. Acesso em: 9 Jun. 2023.
SIEVERT, D. M. et al. Antimicrobial-Resistant Pathogens Associated with Healthcare Associated Infections: Summary of Data Reported to the National Healthcare Safety Network at the Centers for Disease Control and Prevention, 2009–2010. Infection Control and Hospital Epidemiology. v. 34, n. 1, p. 1-14, 2013. Disponível em: <10.1086/668770.>. Acesso em: 9 Jun. 2023.
SILVA, A. S.; FURTADO, S. C.; VARGAS, B. L. Avaliação microbiológica do queijo coalho produzido com leite pasteurizado sob refrigeração. Revista Nanbiquara. [s.l.], v. 6, n. 1, p. 118-133, 2017.
SILVA, N.; IGREJAS, G.; GONÇALVES, A.; POETA, P. Commensal gut bacteria: distribution of Enterococcus species and prevalence of Escherichia coli phylogenetic groups in animals and humans in Portugal. Annals of Microbiology. v. 62, pp. 449-459, 2012. Disponível em: <https://doi.org/10.1016/S1074-5521(02)00198-9>. Acesso em: 9 Jun. 2023.
SILVA, R. A. et al. Can artesanal “coalho” cheese from Northeastern Brazil be used as a functional food? Food Chemistry. v. 135, n. 3, pp. 1533-1538, 2012. Disponível em: <10.1016/j.foodchem.2012.06.058>. Acesso em: 9 Jun. 2023.
SOLACHE, M. G.; RICE, L. B. The Enterococcus: a model of Adaptability to Its Environment. Clinical Microbiology Reviews. v. 32, n. 2, 2019. Disponível em: <10.1128/CMR.00058-18>. Acesso em: 9 Jun. 2023.
SOUZA, A. Z. B. et al. Aspectos físico-químicos e microbiológicos do queijo tipo coalho comercializado em estados do nordeste do Brasil. Arquivos do Instituto Biológico. v. 81, n. 1, p. 30-35, 2014. Disponível em: <10.1590/S1808-16572014000100006>. Acesso em: 9 Jun. 2023.
SURYALETHA, K. et al. Decoding the proteomic changes involved in the biofilm formation of Enterococcus faecalis SK460 to elucidate potential biofilm determinants. BMC Microbiology. v. 19, n. 146, 2019. Disponível em: <https://doi.org/10.1186/s12866-019-1527-2>. Acesso em: 9 Jun. 2023.
TEIXEIRA, I. M. S. Biofilmes e infecções associadas a dispositivos médicos. 2019. Dissertação (Mestrado em Ciências Farmacêuticas), Universidade Fernando Pessoa. Portugal.
TEIXEIRA, L.; MERQUIOR, V. L. Molecular typing in bacterial infections. Humana Press. v. 1, n. 12, p. 17-27, 2013. Disponível em: <10.1007/978-1-62703-185-1_2>. Acesso em: 9 Jun. 2023.
TERÁN, C. G.; RODRÍGUEZ, V. R.; RANGEL, A. C. Analysis of antibiotic uses and resistance in an ICU from Monteria, Colombia. Revista médica Risaralda. v. 24, n. 2, pp. 75-80, 2018. Disponível em: <https://dialnet.unirioja.es/servlet/articulo?codigo=6708347>. Acesso em: 9 Jun. 2023.
TERRA, M. R.; FURLANETO, M. C.; MAIA, L. F. Enterococcus multirresistente a antimicrobianos: um importante patógeno nosocomial. Revista Uningá Review. v. 29, n. 3, p. 94- 102, 2017. Disponível em: <https://www.mastereditora.com.br/periodico/20170304_121005.pdf>. Acesso em: 9 Jun. 2023.
TERRA, M. R.; FURLANETO, M. C.; MAIA, L. FOOD AS A POTENTIAL RESERVOIR OF ENTEROCOCCUS THAT HOST DETERMINANTS OF VIRULENCE AND RESISTANCE. Brazilian Journal of Surgery and Clinical Research. v. 22, n. 1, p. 86-93, 2018. Disponível em: <https://www.mastereditora.com.br/periodico/20180303_175405.pdf>. Acesso em: 9 Jun. 2023.
URQUHART, C. et al. Atividade antibiofilme da cumarina frente à Pseudomonas aeruginosa. Anais do 10º Salão Internacional de Ensino, Pesquisa e Extensão, Brazilian Journal of Development. v. 6, n. 10, pp. 77261–77268, 2019. Disponível em: <https://doi.org/10.34117/bjdv6n10-231>. Acesso em: 9 Jun. 2023.
VALLEJO, M.; PARADA, R. B.; MARGUET, E. R. Aislamiento de cepas de Enterococcus hirae productoras de enterocinas a partir del contenido intestinal de mejillón patagônico (Mytilus edulis platensis). Revista Argentina de Microbiologia. v. 52, n. 2, pp. 136-144, 2019. Disponível em: <https://doi.org/10.1016/j.ram.2019.06.001>. Acesso em: 9 Jun. 2023.
VIGNAROLI, C. et al. Multidrug- resistant Enterococci in meat and faeces and co-transfer of resistance from an Enterococcus durans to a human Enterococcus faecium. Current Microbiology. v. 62, n. 5, p. 1438-1447, 2011. Disponível em: <10.1007/s00284-011-9880-x>. Acesso em: 9 Jun. 2023.
WHO – WORLD HEALTH ORGANIZATION. Zoonoses and the Human-Animal-Ecosystems Interface. [S. l.] WHO, 2017.
[1] Doctoral student of the Graduate Program in Animal Bioscience. ORCID: 0000-0001-9410-7631. Currículo Lattes: http://lattes.cnpq.br/4532231119888940.
[2] PhD in Animal Bioscience. ORCID: 0000-0002-2871-6655. Curriculum Lattes: http://lattes.cnpq.br/3276511555193502.
[3] PhD in Animal Bioscience. ORCID: 0000-0002-1982-7142. Curriculum Lattes:http://lattes.cnpq.br/5753018560783714.
[4] Graduate in Veterinary Medicine. ORCID:0000-0001-9273-5204. Lattes Curriculum: http://lattes.cnpq.br/8329028352662293.
[5] Graduate in Veterinary Medicine. ORCID: 0000-0001-5584-9464. Curriculum Lattes:http://lattes.cnpq.br/1495170820726310.
[6] Doctoral student of the Graduate Program in Animal Bioscience. ORCID: 0000-0003-0698-6125. Curriculum Lattes: http://lattes.cnpq.br/5170743479462841.
[7] PhD in Veterinary Science. ORCID: 0000-0002-0072-5103. Lattes Curriculum: http://lattes.cnpq.br/5676854885081836.
[8] Bachelor’s Degree in Biological Sciences. ORCID: 0000-0001-7149-4931. Lattes Resume: https://lattes.cnpq.br/2928291078391850.
[9] M.Sc. in Animal Bioscience. ORCID: 0000-0002-4913-8313. Curriculum Lattes: http://lattes.cnpq.br/4967459162060058.
[10] PhD in Biochemistry and Physiology, Master in Physiology, Biologist. ORCID: 0000-0003-1493-7964. Curriculum Lattes: http://lattes.cnpq.br/9044747136928972.
[11] Advisor. Doctor of the Graduate Program in Animal Bioscience. ORCID: 0000-0002-1289-2902. Curriculum Lattes: http://lattes.cnpq.br/5998863169551704.
Sent: April 24, 2023.
Approved: June 1, 2023.